Principal Components Analysis of Variance Simultaneous Component Analysis - PC-ANOVA
Source:R/pcanova.R
pcanova.RdThis is a quite general and flexible implementation of PC-ANOVA.
Arguments
- formula
Model formula accepting a single response (block) and predictor names separated by + signs.
- data
The data set to analyse.
- ncomp
The number of components to retain, proportion of variation or default = minimum cross-validation error.
- contrasts
Effect coding: "sum" (default = sum-coding), "weighted", "reference", "treatment".
- ...
Additional parameters for the
hdanovafunction.
Value
A pcanova object containing loadings, scores, explained variances, etc. The object has
associated plotting (pcanova_plots) and result (pcanova_results) functions.
Details
PC-ANOVA works in the opposite order of ASCA. First the response matrix is decomposed using ANOVA. Then the components are analysed using ANOVA with respect to a design or grouping in the data. The latter can be ordinary fixed effects modelling or mixed models.
References
Luciano G, Næs T. Interpreting sensory data by combining principal component analysis and analysis of variance. Food Qual Prefer. 2009;20(3):167-175.
See also
Main methods: asca, apca, limmpca, msca, pcanova, prc and permanova.
Workhorse function underpinning most methods: hdanova.
Extraction of results and plotting: asca_results, asca_plots, pcanova_results and pcanova_plots
Examples
# Load candies data
data(candies)
# Basic PC-ANOVA model with two factors, cross-validated opt. of #components
mod <- pcanova(assessment ~ candy + assessor, data = candies)
print(mod)
#> PC-ANOVA - Principal Components Analysis of Variance
#>
#> Call:
#> pcanova(formula = assessment ~ candy + assessor, data = candies)
# PC-ANOVA model with interaction, minimum 90% explained variance
mod <- pcanova(assessment ~ candy * assessor, data = candies, ncomp = 0.9)
print(mod)
#> PC-ANOVA - Principal Components Analysis of Variance
#>
#> Call:
#> pcanova(formula = assessment ~ candy * assessor, data = candies, ncomp = 0.9)
summary(mod)
#> PC-ANOVA - Principal Components Analysis of Variance
#>
#> Call:
#> pcanova(formula = assessment ~ candy * assessor, data = candies, ncomp = 0.9)
#> $`Comp. 1`
#> Df Sum Sq Mean Sq F value Pr(>F) Error Term
#> candy 4 31470.6052 7867.65131 780.176157 9.969318e-80 Residuals
#> assessor 10 224.9208 22.49208 2.230371 2.089353e-02 Residuals
#> candy:assessor 40 707.7098 17.69275 1.754457 1.158415e-02 Residuals
#> Residuals 110 1109.2900 10.08445 NA NA <NA>
#>
#> $`Comp. 2`
#> Df Sum Sq Mean Sq F value Pr(>F) Error Term
#> candy 4 1573.830 393.45749 33.160399 3.942715e-18 Residuals
#> assessor 10 278.301 27.83010 2.345507 1.502735e-02 Residuals
#> candy:assessor 40 1053.295 26.33238 2.219280 5.888081e-04 Residuals
#> Residuals 110 1305.181 11.86528 NA NA <NA>
#>
#> $`Comp. 3`
#> Df Sum Sq Mean Sq F value Pr(>F) Error Term
#> candy 4 307.1203 76.78008 7.645952 1.790336e-05 Residuals
#> assessor 10 1006.6196 100.66196 10.024169 8.573804e-12 Residuals
#> candy:assessor 40 484.0250 12.10062 1.205010 2.229486e-01 Residuals
#> Residuals 110 1104.6118 10.04193 NA NA <NA>
#>
# Tukey group letters for 'candy' per component
lapply(mod$models, function(x)
mixlm::cld(mixlm::simple.glht(x,
effect = "candy")))
#> $`Comp 1`
#> Tukey's HSD
#> Alpha: 0.05
#>
#> Mean G1 G2
#> 4 11.77480 A
#> 2 11.66764 A
#> 3 10.36430 A
#> 1 -16.83932 B
#> 5 -16.96742 B
#>
#> $`Comp 2`
#> Tukey's HSD
#> Alpha: 0.05
#>
#> Mean G1 G2 G3 G4
#> 5 4.8321239 A
#> 2 1.1912987 C
#> 4 0.3033102 B C
#> 3 -1.9764732 B
#> 1 -4.3502597 D
#>
#> $`Comp 3`
#> Tukey's HSD
#> Alpha: 0.05
#>
#> Mean G1 G2
#> 5 1.2935060 A
#> 3 1.2336431 A
#> 2 0.7696525 A
#> 4 -1.4424540 B
#> 1 -1.8543476 B
#>
# Result plotting
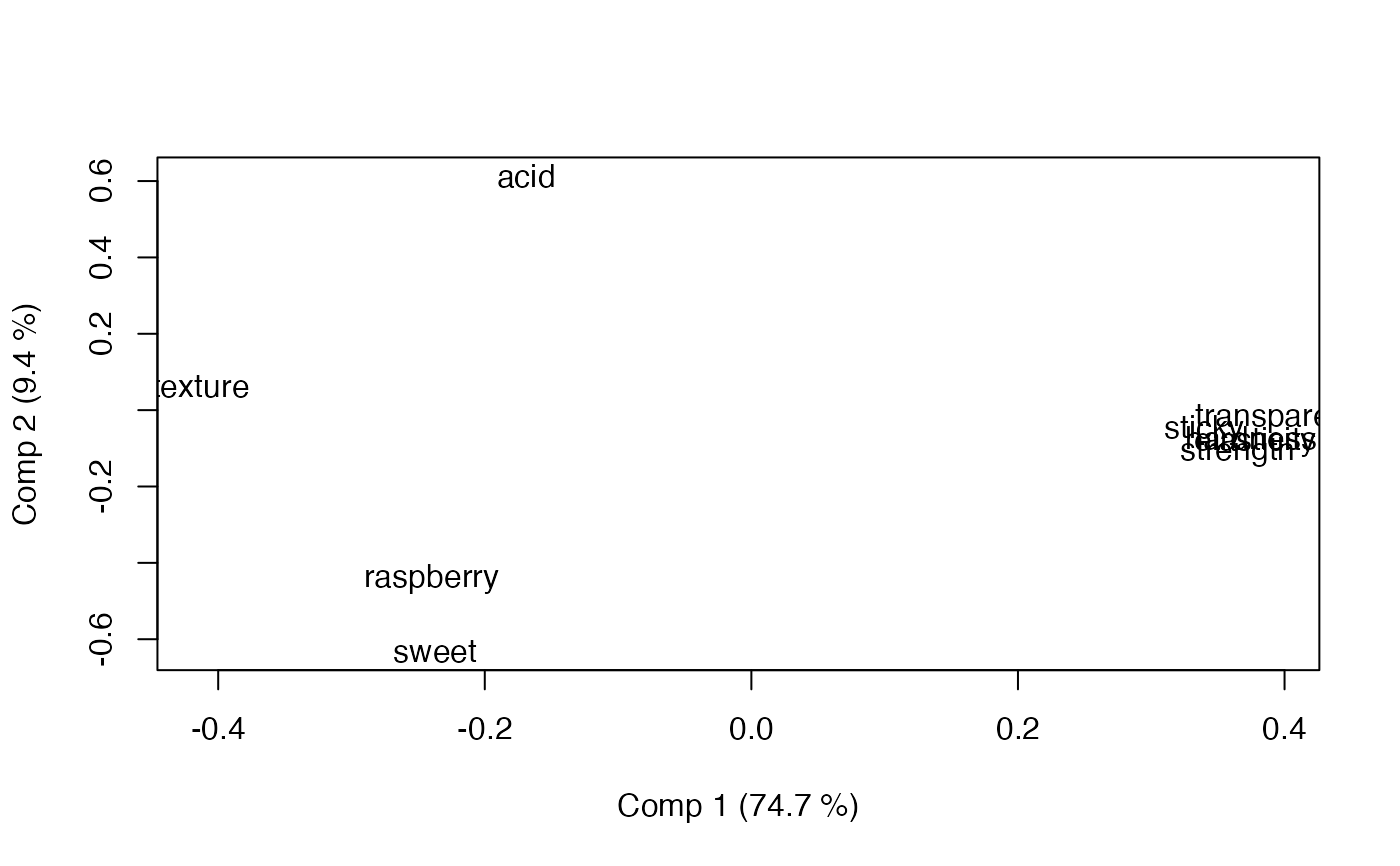
loadingplot(mod, scatter=TRUE, labels="names")
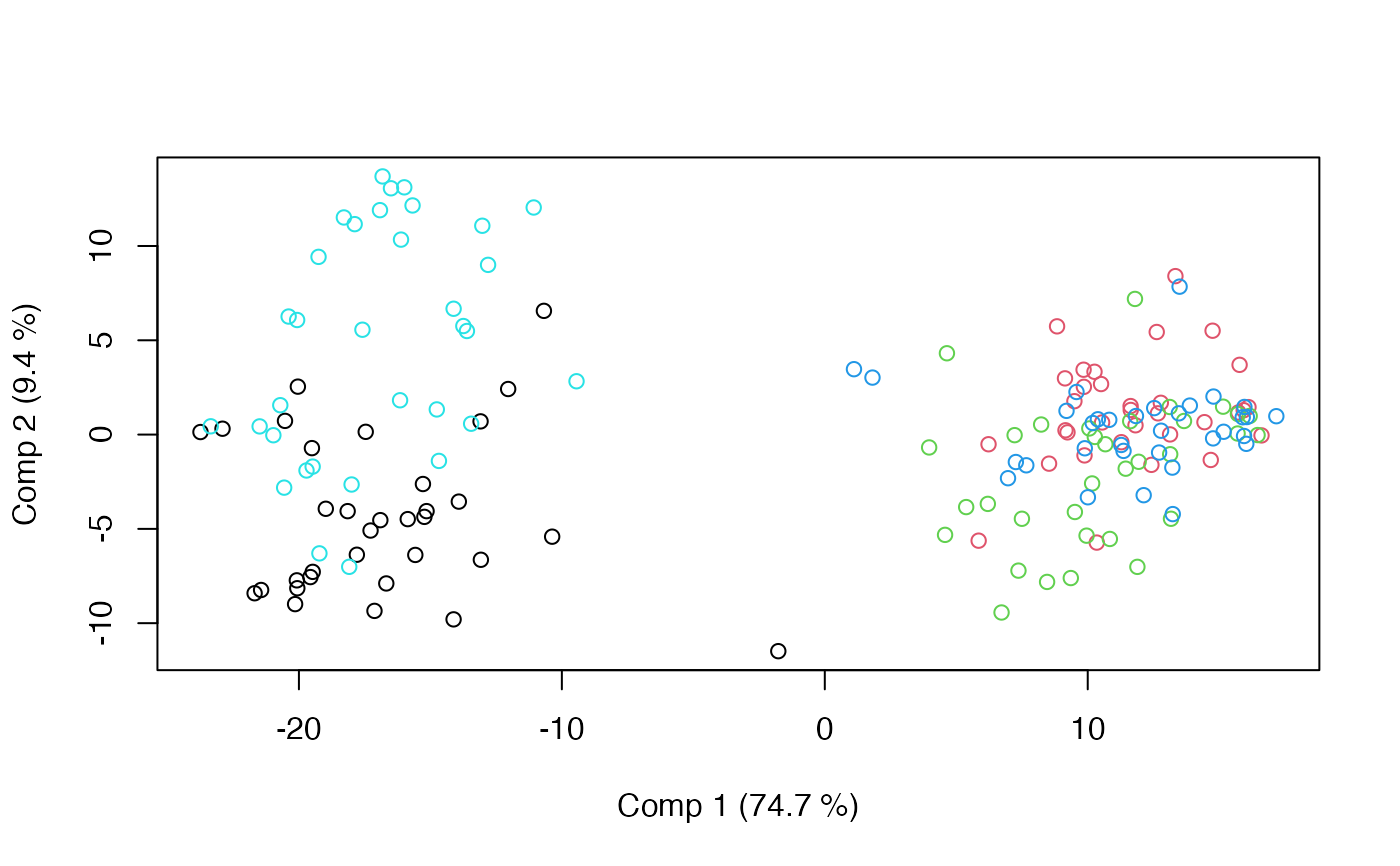
 scoreplot(mod)
scoreplot(mod)
 # Mixed Model PC-ANOVA, random assessor
mod.mix <- pcanova(assessment ~ candy + r(assessor), data=candies, ncomp = 0.9)
scoreplot(mod.mix)
# Fixed effects
summary(mod.mix)
#> PC-ANOVA - Principal Components Analysis of Variance
#>
#> Call:
#> pcanova(formula = assessment ~ candy + r(assessor), data = candies, ncomp = 0.9)
#> $`Comp. 1`
#> Df Sum Sq Mean Sq F value Pr(>F) Error Term
#> candy 4 31470.6052 7867.65131 649.503449 1.371489e-93 Residuals
#> assessor 10 224.9208 22.49208 1.856803 5.555920e-02 Residuals
#> Residuals 150 1816.9999 12.11333 NA NA <NA>
#>
#> $`Comp. 2`
#> Df Sum Sq Mean Sq F value Pr(>F) Error Term
#> candy 4 1573.830 393.45749 25.024047 6.924974e-16 Residuals
#> assessor 10 278.301 27.83010 1.770005 7.067805e-02 Residuals
#> Residuals 150 2358.476 15.72318 NA NA <NA>
#>
#> $`Comp. 3`
#> Df Sum Sq Mean Sq F value Pr(>F) Error Term
#> candy 4 307.1203 76.78008 7.249619 2.301866e-05 Residuals
#> assessor 10 1006.6196 100.66196 9.504560 3.816476e-12 Residuals
#> Residuals 150 1588.6368 10.59091 NA NA <NA>
#>
# Mixed Model PC-ANOVA, random assessor
mod.mix <- pcanova(assessment ~ candy + r(assessor), data=candies, ncomp = 0.9)
scoreplot(mod.mix)
# Fixed effects
summary(mod.mix)
#> PC-ANOVA - Principal Components Analysis of Variance
#>
#> Call:
#> pcanova(formula = assessment ~ candy + r(assessor), data = candies, ncomp = 0.9)
#> $`Comp. 1`
#> Df Sum Sq Mean Sq F value Pr(>F) Error Term
#> candy 4 31470.6052 7867.65131 649.503449 1.371489e-93 Residuals
#> assessor 10 224.9208 22.49208 1.856803 5.555920e-02 Residuals
#> Residuals 150 1816.9999 12.11333 NA NA <NA>
#>
#> $`Comp. 2`
#> Df Sum Sq Mean Sq F value Pr(>F) Error Term
#> candy 4 1573.830 393.45749 25.024047 6.924974e-16 Residuals
#> assessor 10 278.301 27.83010 1.770005 7.067805e-02 Residuals
#> Residuals 150 2358.476 15.72318 NA NA <NA>
#>
#> $`Comp. 3`
#> Df Sum Sq Mean Sq F value Pr(>F) Error Term
#> candy 4 307.1203 76.78008 7.249619 2.301866e-05 Residuals
#> assessor 10 1006.6196 100.66196 9.504560 3.816476e-12 Residuals
#> Residuals 150 1588.6368 10.59091 NA NA <NA>
#>