Function for computing standard SO-PLS based on the interface of the pls package.
Arguments
- formula
Model formula accepting a single response (block) and predictor block names separated by + signs.
- ncomp
Numeric vector of components per block or scalar of overall maximum components.
- max_comps
Maximum total number of components from all blocks combined (<= sum(ncomp)).
- data
The data set to analyse.
- subset
Expression for subsetting the data before modelling.
- na.action
How to handle NAs (no action implemented).
- scale
Logical indicating if variables should be scaled.
- validation
Optional cross-validation strategy "CV" or "LOO".
- sequential
Logical indicating if optimal components are chosen sequentially or globally (default=FALSE).
- segments
Optional number of segments or list of segments for cross-validation. (See
[pls::cvsegments()]).- sel.comp
Character indicating if sequential optimal number of components should be chosen as minimum RMSECV ('opt', default) or by Chi-square test ('chi').
- progress
Logical indicating if a progress bar should be displayed while cross-validating.
- ...
Additional arguments to underlying methods.
Value
An sopls, mvr object with scores, loadings, etc.
associated with printing (sopls_results) and plotting methods (sopls_plots).
Details
SO-PLS is a method which handles two or more input blocks by sequentially performing PLS on blocks against a response and orthogonalising the remaining blocks on the extracted components. Component number optimisation can either be done globally (best combination across blocks) or sequentially (determine for one block, move to next, etc.).
References
Jørgensen K, Mevik BH, Næs T. Combining designed experiments with several blocks of spectroscopic data. Chemometr Intell Lab Syst. 2007;88(2): 154–166.
See also
SO-PLS result functions, sopls_results, SO-PLS plotting functions, sopls_plots, SO-PLS Måge plot, maage, and SO-PLS path-modelling, SO_TDI.
Overviews of available methods, multiblock, and methods organised by main structure: basic, unsupervised, asca, supervised and complex.
Examples
data(potato)
so <- sopls(Sensory ~ Chemical + Compression, data=potato, ncomp=c(10,10),
max_comps=10, validation="CV", segments=10)
summary(so)
#> Data: X dimension: 26 0
#> Y dimension: 26 9
#> Fit method: PKPLS
#> Number of components considered: 10
#>
#> VALIDATION: RMSEP
#> Cross-validated using 10 random segments.
#> 0,0 0,1 0,2 0,3 0,4 0,5 0,6 0,7 0,8 0,9
#> 1.1174 0.9463 0.9470 0.9319 0.9327 1.0831 0.9802 1.0401 1.0562 1.0670
#> 0,10 1,0 1,1 1,2 1,3 1,4 1,5 1,6 1,7 1,8
#> 1.0344 0.8491 0.8039 0.8427 0.7785 0.8005 0.8442 0.8449 0.9155 0.9279
#> 1,9 2,0 2,1 2,2 2,3 2,4 2,5 2,6 2,7 2,8
#> 0.9669 0.7712 0.6732 0.6563 0.6951 0.7161 0.7289 0.7394 0.7929 0.8404
#> 3,0 3,1 3,2 3,3 3,4 3,5 3,6 3,7 4,0 4,1
#> 0.7056 0.6732 0.6582 0.7098 0.7358 0.7529 0.7766 0.8266 0.6964 0.6639
#> 4,2 4,3 4,4 4,5 4,6 5,0 5,1 5,2 5,3 5,4
#> 0.6495 0.6961 0.7294 0.7635 0.8050 0.7057 0.6702 0.6645 0.7261 0.7685
#> 5,5 6,0 6,1 6,2 6,3 6,4 7,0 7,1 7,2 7,3
#> 0.8480 0.7746 0.7380 0.7335 0.7767 0.8225 0.8324 0.7910 0.7826 0.8379
#> 8,0 8,1 8,2 9,0 9,1 10,0
#> 1.0183 0.9750 0.9628 1.0054 0.9663 1.2920
#>
#> TRAINING: % variance explained
#> 0,0 0,1 0,2 0,3 0,4 0,5 0,6 0,7 0,8 0,9
#> X 0 45.87 55.51 62.90 66.46 67.73 74.95 78.26 79.85 82.39
#> ref 0 42.19 54.73 65.01 66.85 73.96 74.10 74.44 77.45 77.57
#> hard 0 39.11 41.97 42.23 43.80 50.95 54.87 56.78 59.50 74.53
#> firm 0 42.55 57.44 59.44 61.06 66.78 68.62 69.02 71.34 76.53
#> elas 0 38.64 65.31 73.51 75.63 77.23 79.25 81.19 83.71 86.57
#> adhes 0 16.13 18.11 26.71 26.74 29.70 42.07 44.57 46.47 46.54
#> grainy 0 23.21 43.78 62.23 64.02 67.18 67.72 69.83 74.79 74.79
#> mealy 0 35.35 41.99 57.36 58.84 67.17 70.33 73.21 77.77 78.88
#> moist 0 24.48 27.71 43.10 44.27 48.41 53.39 62.37 67.60 74.79
#> chewi 0 21.98 22.78 48.17 55.50 59.96 68.39 72.54 76.39 77.42
#> 0,10 1,0 1,1 1,2 1,3 1,4 1,5 1,6 1,7 1,8
#> X 83.18 34.20 64.12 71.71 76.24 78.20 79.49 83.37 84.42 85.59
#> ref 77.57 55.79 64.00 64.25 73.75 77.39 79.53 80.42 80.50 80.96
#> hard 75.43 24.10 41.42 44.05 44.07 44.09 47.61 53.70 53.86 62.46
#> firm 79.31 42.73 54.16 58.42 62.05 63.55 64.91 70.85 70.91 70.93
#> elas 88.14 53.24 59.32 63.95 71.43 74.51 75.33 77.16 78.86 78.88
#> adhes 54.68 17.49 22.70 33.54 33.75 34.70 34.90 39.43 48.51 48.92
#> grainy 74.82 57.69 58.26 58.39 71.89 75.95 76.07 76.72 77.73 79.00
#> mealy 79.25 61.37 65.64 66.70 74.33 75.36 77.44 78.35 79.55 80.15
#> moist 77.22 57.32 58.51 62.12 66.20 66.35 67.13 67.14 69.51 71.02
#> chewi 81.68 53.27 54.25 64.22 66.35 69.59 69.96 72.80 77.64 77.64
#> 1,9 2,0 2,1 2,2 2,3 2,4 2,5 2,6 2,7 2,8
#> X 86.07 42.76 72.61 79.40 81.41 82.69 84.98 88.16 88.87 89.84
#> ref 81.20 76.03 85.17 87.21 89.03 90.67 90.68 91.36 91.36 91.39
#> hard 69.98 25.29 44.10 46.63 46.63 46.88 52.03 61.89 64.48 73.84
#> firm 80.21 55.93 69.19 76.45 76.45 76.80 77.79 82.58 82.64 86.03
#> elas 87.63 63.10 70.21 78.03 78.08 78.17 80.37 82.19 84.54 85.69
#> adhes 49.97 34.43 39.48 46.70 46.78 48.48 54.50 56.16 67.45 72.74
#> grainy 79.50 83.95 84.78 87.19 88.01 88.56 89.28 89.29 89.30 89.49
#> mealy 80.19 81.14 85.67 85.68 87.94 88.28 88.57 89.15 89.69 90.49
#> moist 72.44 67.94 69.06 70.49 72.51 72.53 72.61 72.80 73.61 78.69
#> chewi 78.05 70.55 71.41 77.11 77.11 78.09 78.17 79.79 83.27 86.51
#> 3,0 3,1 3,2 3,3 3,4 3,5 3,6 3,7 4,0 4,1
#> X 63.76 78.59 84.55 86.77 87.73 90.09 91.81 92.46 70.09 80.35
#> ref 84.50 86.31 87.91 89.34 91.72 91.74 93.12 93.13 84.91 87.28
#> hard 29.51 45.43 50.00 50.23 51.56 54.69 61.12 63.12 30.80 49.39
#> firm 62.42 68.64 76.93 76.96 77.84 78.25 82.67 82.72 62.64 73.46
#> elas 69.44 70.75 78.04 78.06 78.07 80.60 82.75 85.36 69.94 73.16
#> adhes 34.51 46.88 50.83 51.04 51.60 57.45 58.26 69.10 64.76 65.66
#> grainy 87.18 87.45 88.87 89.86 90.34 91.06 91.37 91.45 87.20 87.21
#> mealy 84.89 86.14 86.18 88.32 89.25 89.48 91.02 91.95 88.83 88.96
#> moist 70.03 70.05 72.13 74.57 74.71 74.88 74.97 76.40 75.09 76.33
#> chewi 70.56 72.61 77.34 77.39 78.58 78.58 79.89 83.77 82.06 82.29
#> 4,2 4,3 4,4 4,5 4,6 5,0 5,1 5,2 5,3 5,4
#> X 87.12 88.89 89.78 92.03 93.37 75.21 84.27 91.16 92.52 93.47
#> ref 88.13 89.65 92.48 92.54 93.63 85.03 88.23 89.16 90.27 92.85
#> hard 49.95 50.65 50.76 54.96 60.99 41.38 53.88 54.08 54.33 54.70
#> firm 77.00 77.10 77.44 78.23 83.34 68.24 75.97 78.78 78.78 79.40
#> elas 78.98 79.09 79.43 82.78 83.85 72.35 74.56 79.87 80.48 80.91
#> adhes 67.28 68.26 69.21 70.41 70.90 64.77 65.81 67.35 67.83 69.01
#> grainy 89.07 90.46 91.63 92.15 92.30 87.53 87.56 89.67 90.98 91.93
#> mealy 88.99 93.13 94.52 94.64 95.00 89.21 89.65 89.73 93.45 94.51
#> moist 76.60 82.33 82.90 82.90 83.28 79.72 79.91 79.98 83.86 84.08
#> chewi 84.08 84.45 86.24 86.38 86.39 82.11 82.31 83.97 84.76 86.45
#> 5,5 6,0 6,1 6,2 6,3 6,4 7,0 7,1 7,2 7,3
#> X 95.40 77.28 86.22 93.18 94.55 95.63 79.57 88.43 95.46 96.65
#> ref 92.91 86.80 90.43 91.47 92.26 94.07 88.07 91.60 92.62 92.70
#> hard 60.53 45.27 56.94 57.05 57.64 59.71 45.28 56.91 56.99 57.22
#> firm 80.75 68.98 76.56 79.17 79.42 80.55 69.02 76.72 79.16 79.35
#> elas 83.34 72.55 74.96 80.08 80.15 80.26 73.76 76.39 81.36 81.61
#> adhes 70.53 64.77 65.79 67.29 69.17 69.37 65.04 65.96 67.53 68.89
#> grainy 92.38 91.29 91.41 93.80 94.06 94.27 91.78 91.89 94.29 94.34
#> mealy 94.83 90.67 91.24 91.36 94.83 95.44 92.38 92.87 92.98 94.77
#> moist 84.08 82.80 82.90 82.93 85.65 85.67 85.12 85.26 85.29 86.55
#> chewi 86.50 84.60 84.72 86.16 86.49 87.50 86.05 86.22 87.61 87.62
#> 8,0 8,1 8,2 9,0 9,1 10,0
#> X 82.82 88.16 95.96 85.49 90.21 87.08
#> ref 90.82 94.67 94.93 91.52 94.42 91.79
#> hard 52.73 59.90 60.33 52.75 63.37 52.92
#> firm 71.26 80.42 80.49 71.81 80.88 71.82
#> elas 73.78 80.71 81.41 75.22 80.15 76.14
#> adhes 68.88 68.89 69.28 71.74 72.73 74.17
#> grainy 91.80 93.21 94.39 91.99 92.91 92.43
#> mealy 93.59 94.05 94.31 93.75 93.98 93.83
#> moist 86.40 86.60 87.34 87.11 88.40 87.14
#> chewi 87.11 88.40 88.43 87.27 89.53 87.28
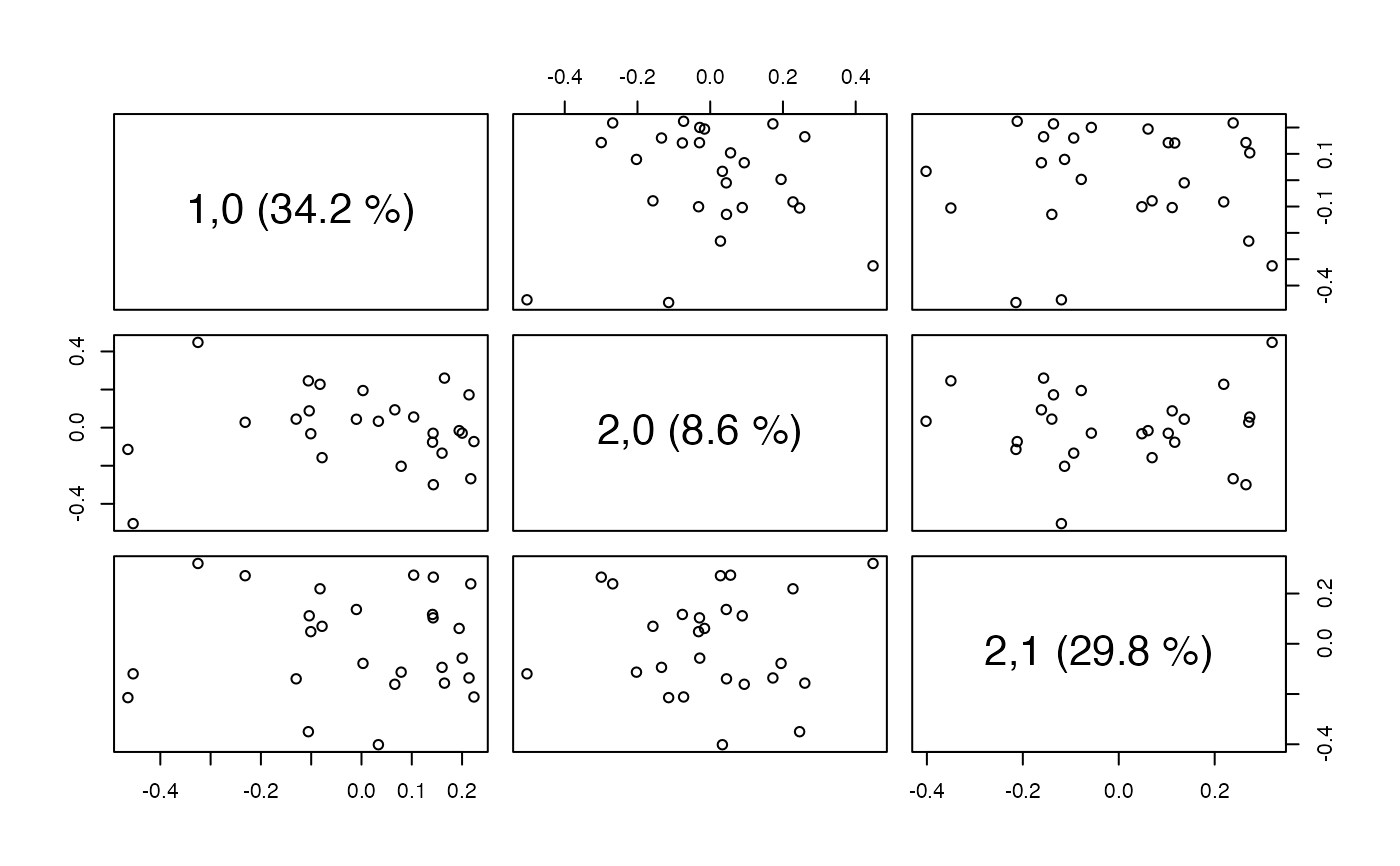
# Scatter plot matrix with two first components from Chemical block
# and 1 component from the Compression block.
scoreplot(so, comps=list(1:2,1), ncomp=2, block=2)
 # Result functions and more plots for SO-PLS
# are found in ?sopls_results and ?sopls_plots.
# Result functions and more plots for SO-PLS
# are found in ?sopls_results and ?sopls_plots.