sMB-PLS is an adaptation of MB-PLS (mbpls) that enforces sparseness in loading weights
when computing PLS components in the global model.
Usage
smbpls(
formula,
data,
subset,
na.action,
X = NULL,
Y = NULL,
ncomp = 1,
scale = FALSE,
shrink = NULL,
truncation = NULL,
trunc.width = 0.95,
blockScale = c("sqrtnvar", "ssq", "none"),
...
)Arguments
- formula
Model formula accepting a single response (block) and predictor block names separated by + signs.
- data
The data set to analyse.
- subset
Expression for subsetting the data before modelling.
- na.action
How to handle NAs (no action implemented).
- X
listof input blocks. If X is supplied, the formula interface is skipped.- Y
matrixof responses.- ncomp
integernumber of PLS components.- scale
logicalfor autoscaling inputs (default = FALSE).- shrink
numericscalar indicating degree of L1-shrinkage/Soft-Thresholding (optional), 0 <= shrink < 1.- truncation
characterindicating type of truncation (optional) "Lenth" uses asymmetric confidence intervals to determine outlying loading weights. "quantile" uses a quantile plot approach to determining outliers.- trunc.width
numericindicating confidence of "Lenth type" confidence interval or quantile in "quantile plot" approach. Default = 0.95.- blockScale
Either a
characterindicating type of block scaling or anumericvector of block weights (see Details).- ...
additional arguments to pls::plsr.
Value
multiblock, mvr object with super-scores, super-loadings, block-scores and block-loading, and the underlying
mvr (PLS) object for the super model, with all its result and plot possibilities. Relevant plotting functions: multiblock_plots
and result functions: multiblock_results.
Details
Two versions of sparseness are supplied: Soft-Threshold PLS, also known as Sparse PLS, and Truncation PLS. The former uses L1 shrinkage of loading weights, while the latter comes in two flavours, both estimating inliers and outliers. The "Lenth" method uses asymmetric confidence intervals around the median of a loading weigh vector to estimate inliers. The "quantile" method uses a quantile plot approach to estimate outliers as deviations from the estimated quantile line. As with ordinary MB-PLS scaled input blocks (1/sqrt(ncol)) are used.
Block weighting is performed after scaling all variables and is by default
"sqrtnvar": 1/sqrt(ncol(X[[i]])) in each block. Alternatives
are "ssq": 1/norm(X[[i]], "F")^2 and "none": 1/1. Finally, if
a numeric vector is supplied, it will be used to scale the blocks
after "ssq" scaling, i.e., Z[[i]] = X[[i]] / norm(X[[i]], "F")^2 * blockScale[i].
References
Sæbø, S.; Almøy, T.; Aarøe, J. & Aastveit, A. ST-PLS: a multi-directional nearest shrunken centroid type classifier via PLS Journal of Chemometrics: A Journal of the Chemometrics Society, Wiley Online Library, 2008, 22, 54-62.
Lê Cao, K.; Rossouw, D.; Robert-Granié, C. & Besse, P. A sparse PLS for variable selection when integrating omics data Statistical applications in genetics and molecular biology, 2008, 7.
Liland, K.; Høy, M.; Martens, H. & Sæbø, S. Distribution based truncation for variable selection in subspace methods for multivariate regression Chemometrics and Intelligent Laboratory Systems, 2013, 122, 103-111.
Karaman, I.; Nørskov, N.; Yde, C.; Hedemann, M.; Knudsen, K. & Kohler, A. Sparse multi-block PLSR for biomarker discovery when integrating data from LC–MS and NMR metabolomics Metabolomics, 2015, 11, 367-379.
See also
Overviews of available methods, multiblock, and methods organised by main structure: basic, unsupervised, asca, supervised and complex.
Examples
data(potato)
# Truncation MB-PLS
# Loading weights inside 60% confidence intervals around the median are set to 0.
tmb <- smbpls(Sensory ~ Chemical+Compression, data=potato, ncomp = 5,
truncation = "Lenth", trunc.width = 0.6)
# Alternative XY-interface
tmb.XY <- smbpls(X=potato[c('Chemical','Compression')], Y=potato[['Sensory']], ncomp = 5,
truncation = "Lenth", trunc.width = 0.6)
identical(tmb, tmb.XY)
#> [1] FALSE
scoreplot(tmb, labels="names") # Exploiting mvr object structure from pls package
 loadingweightplot(tmb, labels="names")
loadingweightplot(tmb, labels="names")
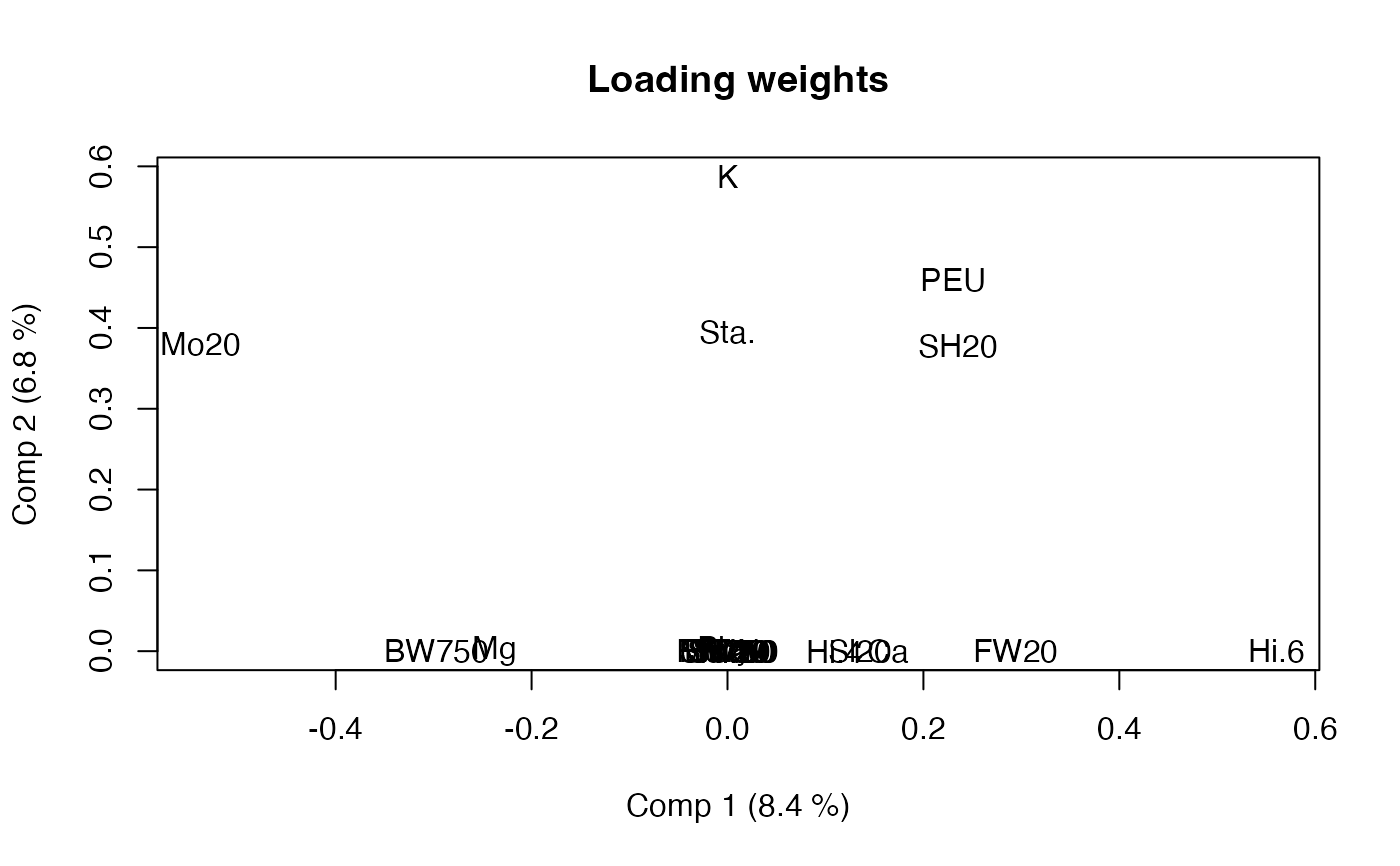 # Soft-Threshold / Sparse MB-PLS
# Loading weights are subtracted by 60% of maximum value.
smb <- smbpls(X=potato[c('Chemical','Compression')], Y=potato[['Sensory']],
ncomp = 5, shrink = 0.6)
print(smb)
#> Sparse Multiblock PLS (Soft-Threshold)
#>
#> Call:
#> smbpls(X = potato[c("Chemical", "Compression")], Y = potato[["Sensory"]], ncomp = 5, shrink = 0.6)
scoreplot(smb, labels="names") # Exploiting mvr object structure from pls package
# Soft-Threshold / Sparse MB-PLS
# Loading weights are subtracted by 60% of maximum value.
smb <- smbpls(X=potato[c('Chemical','Compression')], Y=potato[['Sensory']],
ncomp = 5, shrink = 0.6)
print(smb)
#> Sparse Multiblock PLS (Soft-Threshold)
#>
#> Call:
#> smbpls(X = potato[c("Chemical", "Compression")], Y = potato[["Sensory"]], ncomp = 5, shrink = 0.6)
scoreplot(smb, labels="names") # Exploiting mvr object structure from pls package
 loadingweightplot(smb, labels="names")
loadingweightplot(smb, labels="names")
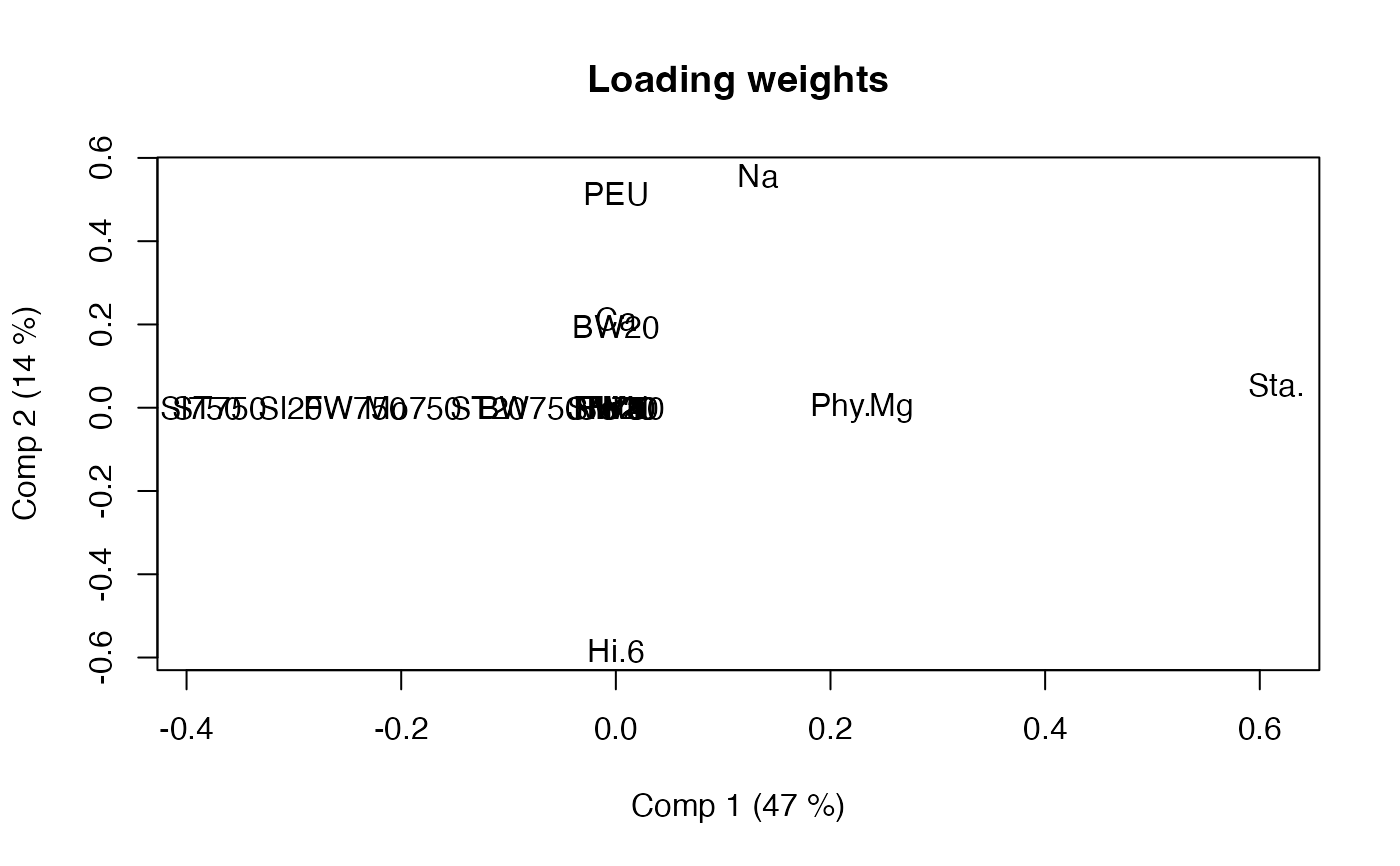 # Emphasis may be different for blocks
smb <- smbpls(X=potato[c('Chemical','Compression')], Y=potato[['Sensory']],
ncomp = 5, shrink = 0.6, blockScale = c(1, 10))
# Emphasis may be different for blocks
smb <- smbpls(X=potato[c('Chemical','Compression')], Y=potato[['Sensory']],
ncomp = 5, shrink = 0.6, blockScale = c(1, 10))