A function computing MB-PLS scores, loadings, etc. on the super-level and block-level.
Usage
mbpls(
formula,
data,
subset,
na.action,
X = NULL,
Y = NULL,
ncomp = 1,
scale = FALSE,
blockScale = c("sqrtnvar", "ssq", "none"),
...
)Arguments
- formula
Model formula accepting a single response (block) and predictor block names separated by + signs.
- data
The data set to analyse.
- subset
Expression for subsetting the data before modelling.
- na.action
How to handle NAs (no action implemented).
- X
listof input blocks. If X is supplied, the formula interface is skipped.- Y
matrixof responses.- ncomp
integernumber of PLS components.- scale
logicalfor autoscaling inputs (default = FALSE).- blockScale
Either a
characterindicating type of block scaling or anumericvector of block weights (see Details).- ...
additional arguments to pls::plsr.
Value
multiblock, mvr object with super-scores, super-loadings, block-scores and block-loading, and the underlying
mvr (PLS) object for the super model, with all its result and plot possibilities. Relevant plotting functions: multiblock_plots
and result functions: multiblock_results.
Details
MB-PLS is the prototypical component based supervised multiblock method.
It was originally formulated as a two-level method with a block-level and a super-level,
but it was later discovered that it could be expressed as an ordinary PLS on concatenated
weighted X blocks followed by a simple loop for calculating block-level loading weights,
loadings and scores. This implementation uses the plsr function on the
scaled input blocks (1/sqrt(ncol)) enabling all summaries and plots from the pls
package.
Block weighting is performed after scaling all variables and is by default
"sqrtnvar": 1/sqrt(ncol(X[[i]])) in each block. Alternatives
are "ssq": 1/norm(X[[i]], "F")^2 and "none": 1/1. Finally, if
a numeric vector is supplied, it will be used to scale the blocks
after "ssq" scaling, i.e., Z[[i]] = X[[i]] / norm(X[[i]], "F")^2 * blockScale[i].
References
Wangen, L.E. and Kowalski, B.R. (1988). A multiblock partial least squares algorithm for investigating complex chemical systems. Journal of Chemometrics, 3, 3–20.
Westerhuis, J.A., Kourti, T., and MacGregor,J.F. (1998). Analysis of multiblock and hierarchical PCA and PLS models. Journal of Chemometrics, 12, 301–321.
See also
Overviews of available methods, multiblock, and methods organised by main structure: basic, unsupervised, asca, supervised and complex.
Examples
data(potato)
# Formula interface
mb <- mbpls(Sensory ~ Chemical+Compression, data=potato, ncomp = 5)
# ... or X and Y
mb.XY <- mbpls(X=potato[c('Chemical','Compression')], Y=potato[['Sensory']], ncomp = 5)
identical(mb$scores, mb.XY$scores)
#> [1] TRUE
print(mb)
#> Multiblock PLS
#>
#> Call:
#> mbpls(formula = Sensory ~ Chemical + Compression, data = potato, ncomp = 5)
scoreplot(mb, labels="names") # Exploiting mvr object structure from pls package
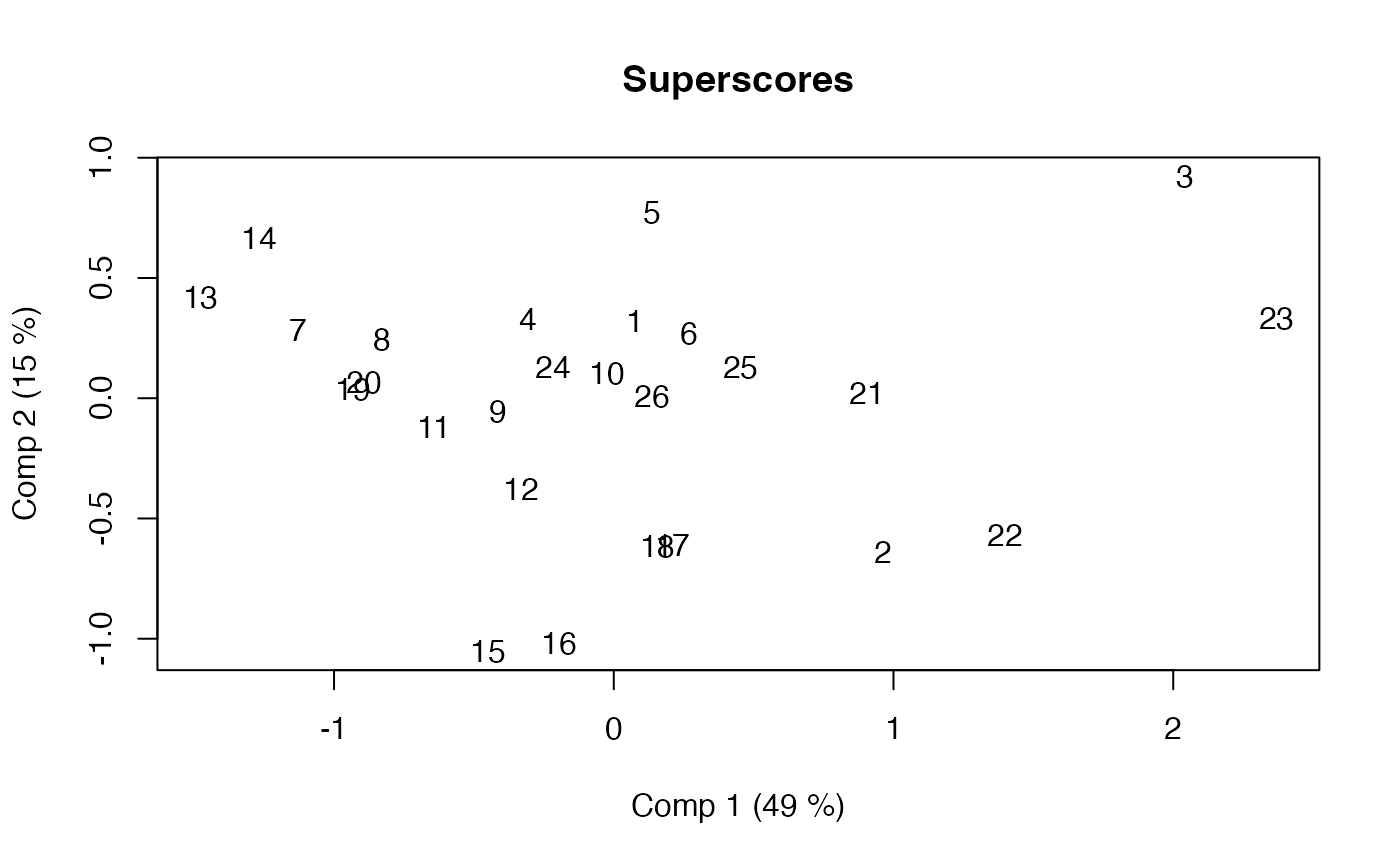 # Block scaling with emphasis on first block
mbs <- mbpls(Sensory ~ Chemical+Compression, data=potato, ncomp = 5, blockScale = c(10, 1))
scoreplot(mbs, labels="names") # Exploiting mvr object structure from pls package
# Block scaling with emphasis on first block
mbs <- mbpls(Sensory ~ Chemical+Compression, data=potato, ncomp = 5, blockScale = c(10, 1))
scoreplot(mbs, labels="names") # Exploiting mvr object structure from pls package
