Rotation testing for HDANOVA. This function performs
random orthogonal rotations of exchangeable residual units for each approved
effect and adds the resulting null distributions to the hdanova object.
Usage
rotation(
object,
rotate = 1000,
unique.digits = 12,
unique.frac = 0.95,
block.type = c("denominator", "global")
)Arguments
- object
A
hdanovaobject.- rotate
Number of random rotations to perform (default = 1000).
- unique.digits
Number of digits used when rounding rotation SSQ values before checking uniqueness (default = 12). Set to
NULLto disable this warning.- unique.frac
Minimum fraction of unique rounded SSQ values required to avoid warning (default = 0.95). Set to
NULLto disable this warning.- block.type
Rotation blocking strategy.
"denominator"(default) rotates within denominator-compatible exchangeable blocks."global"rotates across all observations.
Value
An updated hdanova object with rotation-test results stored in
object$permute for compatibility with existing summary and plotting tools.
See also
Base fitting: hdanova.
Permutation alternative: permutation.
Plot helper: rotationplot.
Examples
# Load candies data
data(candies)
# Basic HDANOVA model with two factors
mod <- hdanova(assessment ~ candy + assessor, data=candies)
# Rotation test
modRot <- rotation(mod)
summary(modRot)
#> High-Dimensional Analysis of Variance fitted using 'lm' (Linear Model)
#> - SS type II, sum coding, restricted model, least squares estimation, SSQ method: qr_regression, 1000 rotations
#> Sum.Sq. Expl.var.(%) p-value
#> candy 33416.66 74.48 0
#> assessor 1961.37 4.37 0
#> Residuals 9489.25 21.15 NA
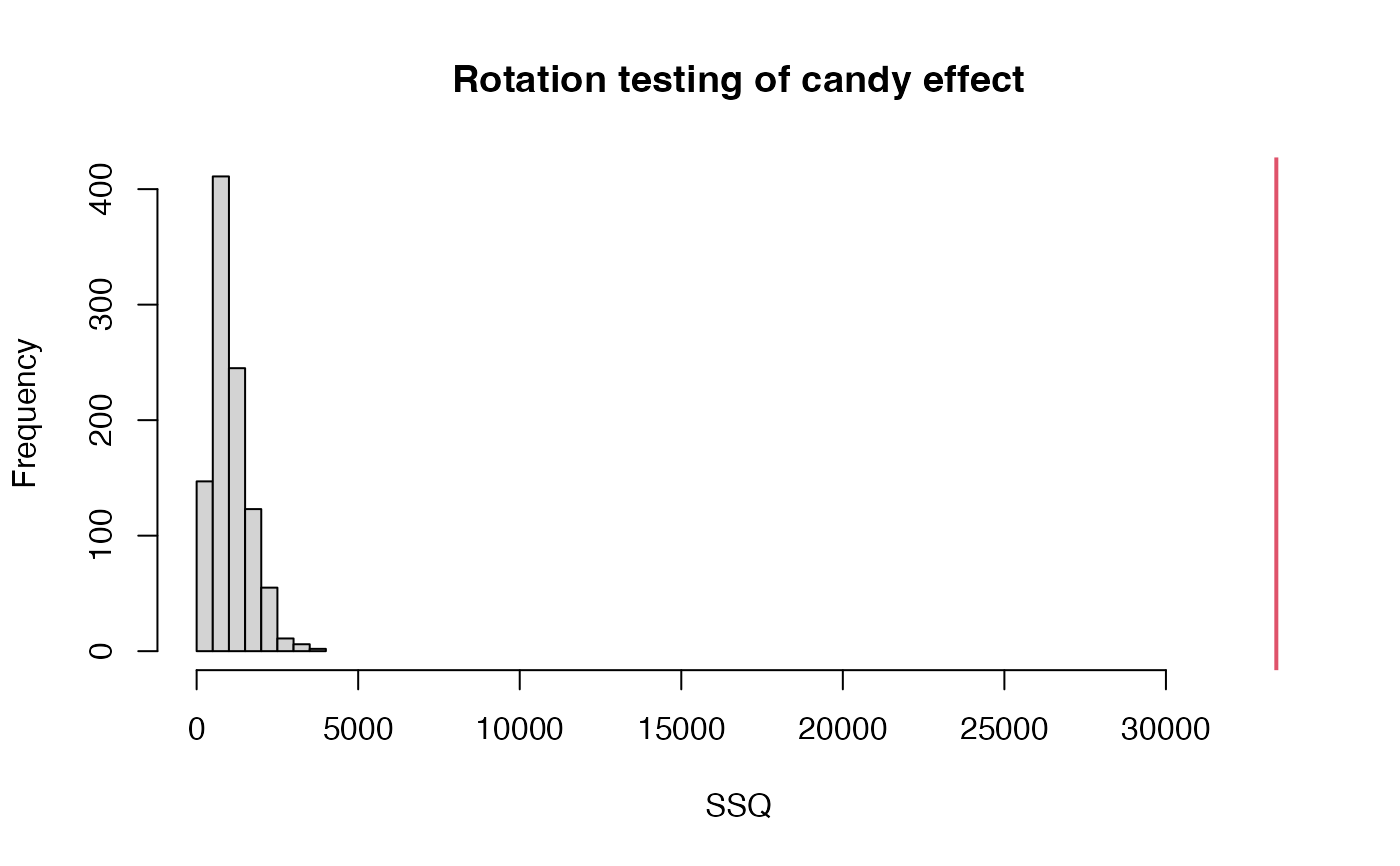
# Plot null distribution for "candy" effect
rotationplot(modRot, factor="candy")